Every year, the effectiveness of the flu vaccine becomes a topic of conversation. While flu season is behind us this year, some burning questions still remain. Why do some people who receive the vaccine get protection from becoming ill while others are hit with the classic chills, nausea, and fever that we all associate with the flu? Are there ways in which we can predict whether or not someone will respond appropriately to the vaccine? Adriana Tomic and colleagues at Stanford University and the University of Oxford are interested in these questions. The answers may lie within a computer program. Published earlier this summer in The Journal of Immunology, Tomic et al. developed an automated machine learning program that could separate high versus low responders to the flu vaccine based on a set of immune system parameters. Through this approach, they identified subsets of immune cells not previously described to have a role in immunity to the influenza virus.
Automated machine learning is considered a subset of artificial intelligence in which a computer is trained to complete a specific task. The best tasks are those that classify items or predict outcomes with defined parameters. For example, say you are a medical doctor interested in improving the diagnosis of a disease by X-ray. You show the computer program a set of X-rays which you tag as being indicative of disease or not. This is considered the ‘training set’ of data. The larger and more diverse the training set is, the more accurate the computer program becomes at completing the specific task. To validate the computer program’s ability to distinguish disease via X-ray, you would provide the computer with X-rays it has not yet seen, a ‘test set’, and analyze its ability to appropriately classify the X-ray. The accuracy, specificity, and sensitivity of the machine learning model can be determined by how often the program is correct at identifying disease or at identifying absence of disease. There are many different machine learning algorithms that suit different types of data distributions, and individual algorithms must be tested for their suitability to complete the task at hand. This technology continues to be applied in the field of medicine, molecular biology, and even in the development of self-driving vehicles.
You may ask yourself, how can this type of program be useful in the field of immunology? The immune system is extremely complex and responses differ according to cell type, time scale, and area of the body. It is difficult to design experimental studies which have large sample sizes, focus on multiple parameters, address variation between different populations of individuals, and capture the entire immune response. Additionally, large immunologic data sets can be difficult to analyze, creating the potential for introduced bias. As a result, immunologic studies are often inconsistent, leading to missing pieces in the puzzle of vaccine response. Tomic et al. comment on the difficulty in working with clinical data, which often spans across multiple years and studies. Due to the experimental capacity at the time of a study, one clinical laboratory or hospital may have different types of data for different patients. For example, patient blood samples may have been evaluated for level of circulating antibody in 2007, but analyzed for immune cell subsets in 2014. Comparing these two types of data can be like comparing apples and oranges, making it tricky to incorporate clinical data into large scale analyses.
Tomic et al. designed their machine learning pipeline, Sequential Iterative Modeling “OverNight,” or SIMON for short, to account for some of these challenges and to more accurately predict flu vaccine responses. SIMON incorporates an algorithm to generate smaller subsets of data from larger clinical databases based on shared immunologic features. This reduces the amount of introduced bias and allows for researchers to utilize original datasets intact. In this study, researchers incorporated 3800 parameters including immune cell subtypes, serum analytes, protein phosphorylation status, and cell signaling capacity from 187 individuals. The large breadth of parameters represented in this study allowed for researchers to get both a cellular and whole body view of the response to the flu vaccine. Each generated dataset was separated into a training set and a test set of values. Then, 128 different machine learning algorithms were applied to train the computer system to identify immunologic patterns that could accurately categorize flu vaccine responders. The most appropriate models were determined based on their ability to correctly identify high versus low responders. Antibody titers, which measure how much antibody specific against the influenza virus a person has circulating in his or her blood, were measured on the day of vaccine administration and 28 days later. If the amount of antibody detected at 28 days was at least four times greater than the amount detected when the vaccine was administered, this was considered a high response. Antibody titers are used as the comparison measure for whether the machine learning algorithms correctly identified responders.
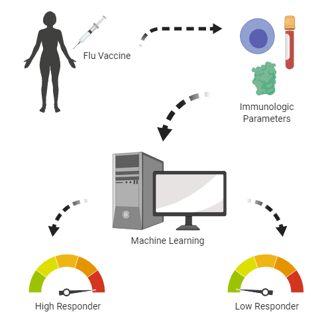
Tomic et al. designed a machine learning computer algorithm which successfully categorized individuals as high or low responders to the flu vaccine based on a set of immunologic parameters. Image produced by Dilara Kiran using Biorender.
Not only was SIMON able to correctly classify high and low flu vaccine responders, it identified unique immunologic features of those populations. Specifically, T cell subtypes not previously thought to play a role in influenza antibody production were demonstrated to be more prevalent in high responders. This was validated experimentally in cell culture by the research group. SIMON streamlined an algorithm selection process that can often be labor intensive, and identified novel immune cell subsets correlated to vaccine response. These cell types could be targeted in future vaccine formulations to increase the protective capacity of the vaccine.
This computational approach to immunology opens doors for researchers to ask new questions and analyze data in ways that were not possible decades ago. As our understanding of the immune system improves and as our technological capacities increase, larger quantities of data will be generated and new analytical tools will need to be created to accommodate data expansion. The machine learning approach is not yet perfect and there are concerns about computer access to personal health data and over reliance of doctors on computer programs to make medical decisions. However, this study demonstrates the importance of applying multidisciplinary approaches to solving health problems. Embracing technological advances and merging them with a better understanding of how our bodies respond to disease can help facilitate the development of better preventative measures, like vaccines.
Dilara Kiran is in her sixth year of the Combined Degree DVM/PhD program at Colorado State University. She aspires to use her knowledge of both clinical practice and research to contribute
to evidence-based scientific policy and is passionate about science communication. Follow her on Twitter @dvmphd2be.
Primary research article: Tomic, A., Tomic, I., Rosenberg-Hasson, Y., Dekker, C.L., Maecker, H.T., and Davis, M.M. (2019). SIMON, an Automated Machine Learning System, Reveals Immune Signatures of Influenza Vaccine Responses. J Immunol 203, 749–759.
Additional References:
Brynjolfsson, E., and Mitchell, T. (2017). What can machine learning do? Workforce implications. Science 358, 1530–1534.
Butler, K.T., Davies, D.W., Cartwright, H., Isayev, O., and Walsh, A. (2018). Machine learning for molecular and materials science. Nature 559, 547.
Shalev-Shwartz, S., and Ben-David, S. (2014). Understanding Machine Learning: From Theory to Algorithms.
Topol, E.J. (2019). High-performance medicine: the convergence of human and artificial intelligence. Nature Medicine 25, 44.